 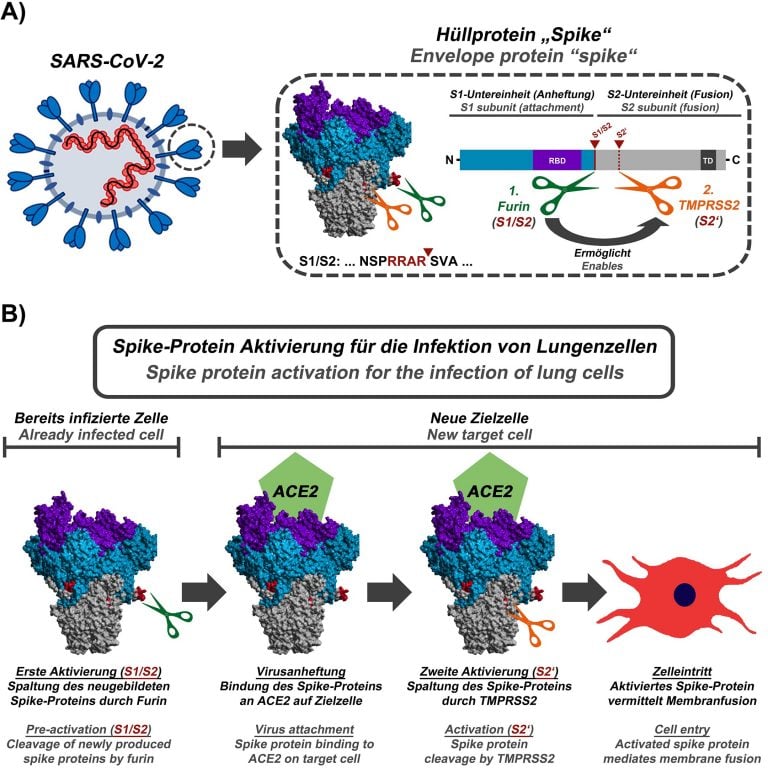
SARS-CoV-2 receptor-binding motif has a more pronounced receptor-interacting ridge and stabilizing residues that increase the affinity for ACE2 compared with SARS-CoV-1 27. These structures revealed that ACE2 interacts with the receptor-binding motif, approximately composed of residues 438–508 within the RBD 26. Structures of the isolated RBD alone and trimeric spike in complex with ACE2 were solved shortly thereafter 16, 22, 23, 24, 25. Subsequent structures mapped out the dynamic nature of the three RBD domains (Fig. 1b) and in another view, one RBD was pointing up (partially open state) 20. In one view, the three RBDs were all pointing down (closed state, Fig. Early structures provided a glimpse of the prefusion complex in two conformations. Spike is the first structural SARS-CoV-2 protein to be deposited in the protein data bank 19. A second cleavage within the S2 subdomain may further enhance infectivity via exposure of the fusion peptide (FP) necessary for viral entry, similar to SARS-CoV-1 17, 18. Furthermore, cleavage at the furin site increases SARS-CoV-2 infectivity 12, 15, 16. Membrane fusion is enhanced by spike cleavage at the furin site separating the S1 and S2 domains and further increasing RBD-open states 12, 13, 14. ACE2 is recognized by the spike receptor-binding domain (RBD) in the up conformation, located within the S1 subunit of the spike protein. Spike mediates host cell entry by binding the cell angiotensin-converting enzyme 2 (ACE2) receptor and by subsequent virion–host membrane fusion. Each virion is decorated with an average of 25–50 spike trimers 7, 8, 9, 10 per virion, although other coronaviruses possess ~90 trimers 11. Spike is a transmembrane homotrimeric class I fusion protein. Spike is the most immunogenic of these proteins in other coronaviruses and is thus a major focus for vaccine, therapeutic and diagnostic development 4, 5, 6. Mature virions include spike (S), the nucleocapsid protein (N), an ion channel (E) and an integral membrane protein (M). Viral RNA is initially translated into two polyproteins with subsequent expression of multiple subgenomic mRNAs that are further processed into smaller proteins by virally encoded proteases 3. 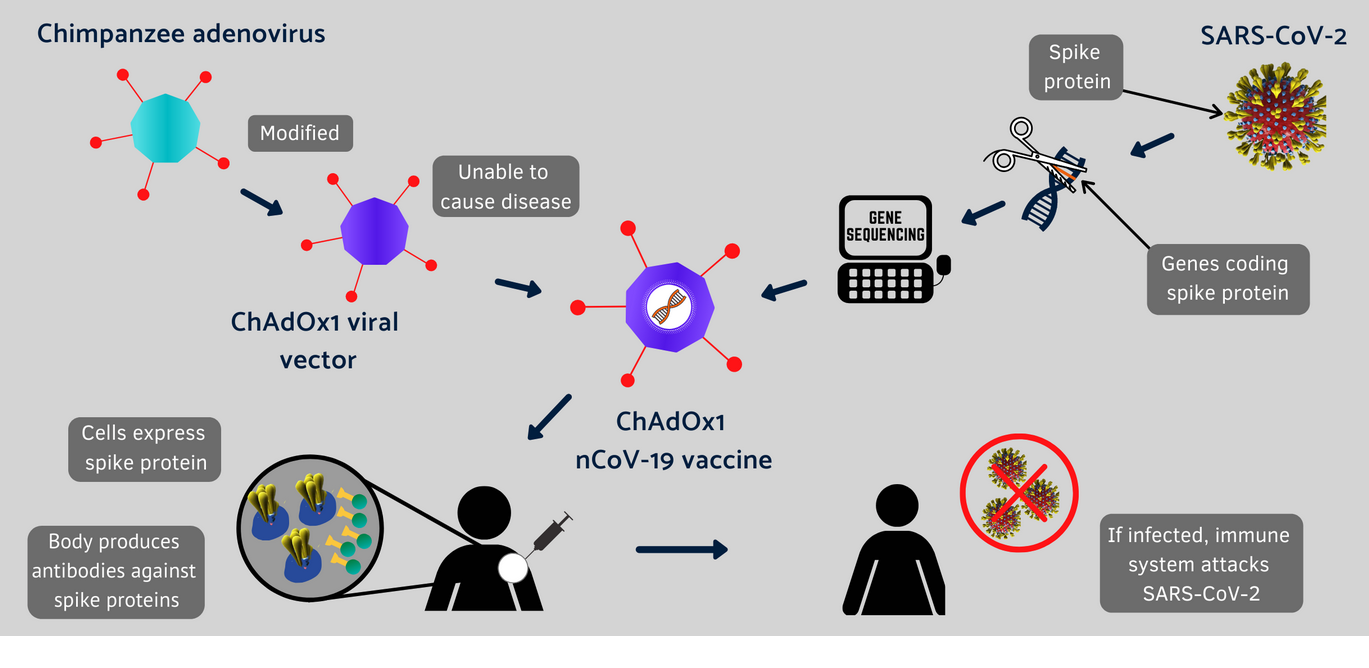
SARS-CoV-2 encodes at least 12 canonical open reading frames in its ~29.9 kilobase RNA genome 1, 2. The causative agent of COVID-19 is severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), a coronavirus-family RNA virus. The coronavirus disease 2019 (COVID-19) pandemic is a global health emergency that has resulted in over four million deaths as of summer 2021. The entire protocol, from transfection to biophysical characterization, can be completed in 7 d by researchers with basic tissue cell culture and protein purification expertise. We also provide guidance on expression quality control, long-term storage, and uses in enzyme-linked immunosorbent assays. Although we focus on HexaPro, this protocol has been used to purify over a hundred different spike variants in our laboratories. This protocol describes an optimized workflow for expressing and biophysically characterizing rationally engineered spike proteins in Freestyle 293 and ExpiCHO cell lines. We recently developed a prefusion-stabilized spike variant, termed HexaPro for six stabilizing proline substitutions, that can be expressed with a yield of >30 mg/L in ExpiCHO cells. 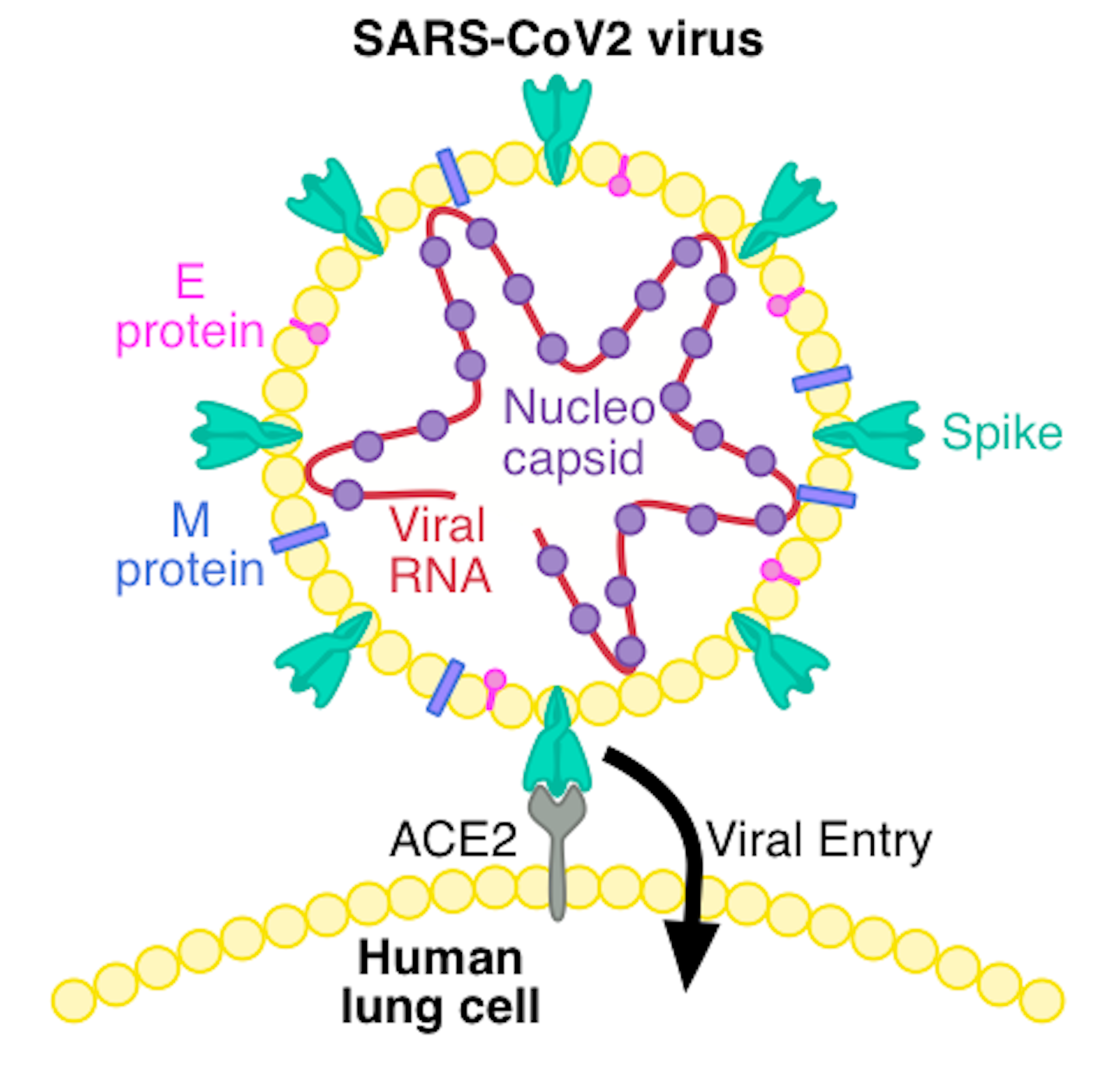
However, the spike protein is difficult to produce recombinantly because it is a large trimeric class I fusion membrane protein that is metastable and heavily glycosylated. The severe acute respiratory syndrome coronavirus 2 spike protein is a critical component of coronavirus disease 2019 vaccines and diagnostics and is also a therapeutic target. Nature Protocols volume 16, pages 5339–5356 ( 2021) Cite this article Expression and characterization of SARS-CoV-2 spike proteins 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
